Title
Seeing Science Clearly
Subhead
New tools from the Coskun Lab help evaluate biomedical data visualizations.
ID
Mar 11, 2026
| By Leeanna Allen
News Image

Image Caption
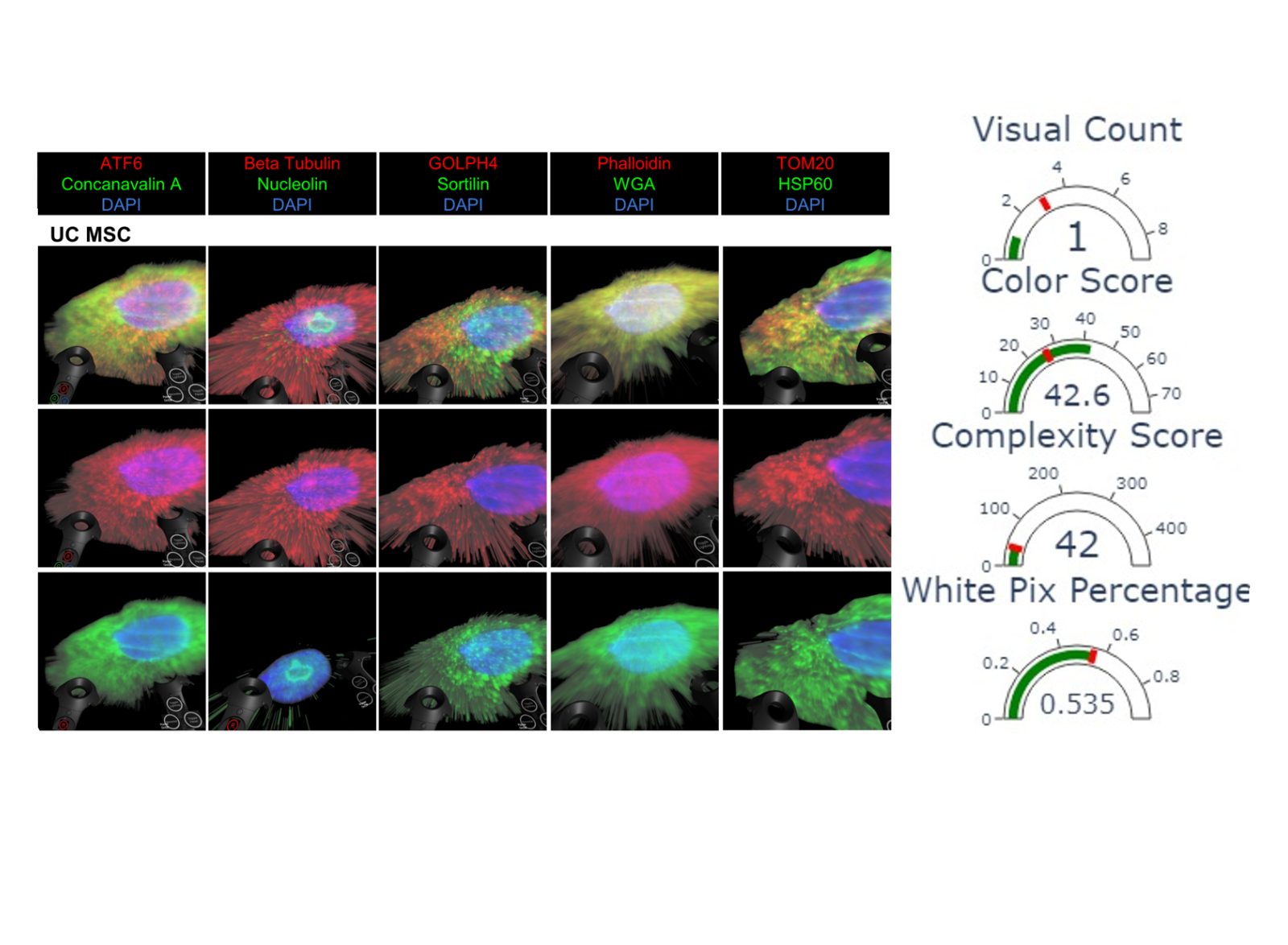
A new algorithm helps biomedical engineers determine the quality of scientific figures like this one.