Dr. Coskun is a Bernie-Marcus Early-Career Professor of Biomedical Engineering at Georgia Institute of Technology and Emory University. Dr. Coskun is a systems biotechnologist and bioengineer, working at the nexus of multiplexed cell imaging and quantitative tissue biology. Dr. Coskun directs an interdisciplinary research team at the Single Cell Biotechnology and Spatial Omics Laboratory, an interdisciplinary program strategically positioned for multiparameter imaging one cell at a time by spatial context and function. Dr. Coskun holds 5 issued patents and is also the co-author of more than 50 peer-reviewed publications in major scientific journals. He is a recipient of the NSF CAREER Award 2024, NIH R35 MIRA Award 2023, Sigma Xi Young Faculty Award 2025, CMBE Young Innovator Award 2024, BMES-CMBE Rising Star Award 2023, American Lung Association Innovation Award 2022, Burroughs Welcome Fund CASI Award 2016, and Student Recognition of Excellence in Teaching: Class of 1934 CIOS Award, among other research and teaching awards. Previously, Dr. Coskun was an Instructor at Stanford University. Dr. Coskun received his postdoctoral training from the California Institute of Technology. He holds a Ph.D. degree from the University of California, Los Angeles.His research has been supported by Federal and Private grants, including the National Institutes of Health (NIGMS, NIA, NIAID, NCI, NIDCR, OD, and ORIP), Wellcome LEAP, Burroughs Wellcome Fund (CASI), NSF CMaT, American Cancer Society IRG, Multi-cellular engineered living systems (M-CELS), and Regenerative Medicine Center. In addition, he leads outreach programs to engage K12 students and undergraduate students through BioCrowd Studio, an innovative crowd-sourcing program bringing together interactive virtual media, distributed biokits, and collaborative spatial discovery.
Coulter BME Professor Earns Tenure, Eyes Future of Innovation in Health and Medicine
Coskun pioneering new research area and building a company around iseqPLA technology
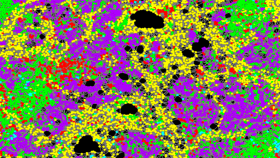
Biomedical engineer will present groundbreaking mapping tool aimed at drug resistant cancers at BMES Annual Meeting
BME researcher Yue Chen using NSF CAREER Award to develop MRI-safe surgical robot
Microscale Single Cell Cancer Metabolism Research Earns Honor for Coulter BME Assistant Professor